AMD Projects: Improving TB Detection and Surveillance
Using whole genome sequencing to transform tuberculosis outbreak detection and surveillance
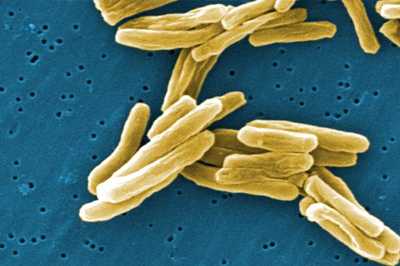
An outbreak investigation aims to stop transmission of TB. This public health intervention involves rapidly identifying and treating all infectious persons.
Outbreaks of Mycobacterium tuberculosis (TB) continue to be a challenge in the United States. Public health professionals use laboratory methods, such as genotyping, to determine when people get TB from the same strain of bacteria. Genotyping can help scientists pinpoint outbreaks that otherwise might be hidden. Since TB that spreads from one person to another will have the same genetic make-up, scientists can isolate bacteria from different people in the laboratory and compare the genetic sequences to find cases caused by the same strain of the bacteria.
But this tool also can mistakenly indicate an outbreak where there isn’t one, especially for genotypes that are common in the population. False matches can slow down the work of local public health professionals as they try to identify, track, and control a potential outbreak when this occurs. It also consumes resources without leading to an effective solution.
CDC scientists studying TB believe genomic surveillance has the power to increase the accuracy of outbreak detection systems. Whole genome sequencing (WGS) lets CDC scientists examine about 90% of the TB genome, allowing them to measure the genetic differences between isolates more easily. CDC expects that the data generated through genomic surveillance will provide more accurate information to public health officials.
Through the work of this project, CDC hopes to implement genomic surveillance at the national level and build the capacity of state laboratory professionals to conduct WGS for TB. In addition, this work has the potential to help local public health officials more precisely identify how the infection has spread and which locations are most important to field investigations in an outbreak. This will ensure interventions have the greatest impact and protect people’s health even better. At the same time, genomic surveillance will be more cost efficient by focusing investigations on cases that are truly linked.
2017 Project Update
Since the project began in 2014, investigators have used advanced molecular detection (AMD) technologies to characterize thousands of Mycobacterium tuberculosis isolates from patients in more than 80 genotype clusters nationally. This information has allowed local and state TB control programs to focus resources on those clusters and patients with recent TB transmission where interventions are most useful.
Investigators also sequenced approximately 5,000 M. tuberculosis isolates to create a new analytic pipeline. While the previous analytic pipelines only allowed for the comparison of relatively small numbers of similar strains, this new pipeline allows for comparison of thousands of diverse strains. Additionally, to help expand sequencing capacity, CDC funded five state public health laboratories in 2016 to conduct WGS on approximately 200 M. tuberculosis isolates per month. As work continues, CDC is getting closer to the ultimate goal of developing the expertise, tools, and infrastructure necessary nationally to conduct genomic surveillance of all isolates of M. tuberculosis in the United States.
- Page last reviewed: April 13, 2016
- Page last updated: April 13, 2016
- Content source:


 ShareCompartir
ShareCompartir