AMD Projects: Putting Flu Surveillance in the Cloud
Using advanced methods to track ever-changing flu viruses
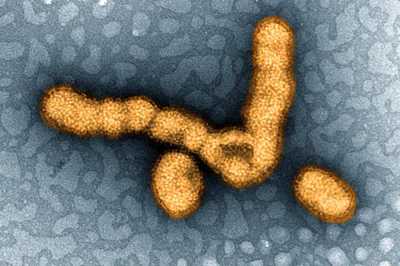
CDC is strengthening influenza surveillance by building capacity of state labs to use AMD methods to obtain more data on flu viruses, such as H1N1.
Influenza (flu) viruses are constantly changing. They can change from one season to the next and even within the course of one flu season. These changes can result in the seasonal flu vaccine providing less than optimal protection or in the emergence of new influenza viruses against which people have no preexisting immunity. For example, a new H1N1 flu virus emerged in 2009 after the seasonal vaccine was formulated. Though millions of people received the seasonal flu vaccine, it did not protect them from this particular H1N1 strain, which spread in people and caused a global flu pandemic.
Year round, scientists from CDC, World Health Organization (WHO), and other partners monitor the influenza viruses that are spreading among people. These scientists study the viruses in the laboratory to track how they are changing. The U.S. influenza surveillance network includes 64 state and local public health laboratories that routinely test patient samples for influenza. Once these labs confirm influenza, they send a subset of specimens to CDC for deeper analysis. CDC is able to use advanced molecular detection (AMD) to confirm the state’s testing results, examine viruses in greater detail using next-generation genomic sequencing (NGS), and then perform complex computing to reveal the precise make-up of the virus’ genes.
In this project, CDC is laying the groundwork to help state public health laboratories develop the capacity to use AMD on influenza viruses. CDC is partnering with Wisconsin State Laboratory of Hygiene (WSLH) and Association of Public Health Laboratories (APHL) to demonstrate that state public laboratories can use NGS to obtain the same results as CDC’s labs. CDC and WSLH are also developing an APHL-hosted cloud-based mechanism for assembling and analyzing influenza sequences generated through the WSLH NGS effort. Ultimately, the lessons learned and tools developed through the partnership will serve as a model that can be applied to other state public health labs and to other pathogens.
- Page last reviewed: October 16, 2015
- Page last updated: October 16, 2015
- Content source:


 ShareCompartir
ShareCompartir