Ambidensovirus
The virus genus Ambidensovirus[1] belongs to the Densovirinae subfamily[2] of the family Parvoviridae.[3][4] Members of this genus are single-stranded DNA viruses. This genus infects invertebrates, including insects, crustacea and echinoderms. There are currently eleven species in this genus including the type species Lepidopteran ambidensovirus 1.[5][6]
| Ambidensovirus | |
|---|---|
| Virus classification | |
| Group: | Group II (ssDNA) |
| Order: | Unassigned |
| Family: | |
| Subfamily: | Densovirinae |
| Genus: | Ambidensovirus |
| Type species | |
| Lepidopteran ambidensovirus 1 | |
| Species | |
| |
Taxonomy
Group: ssDNA
- Family: Parvoviridae
- Sub-Family: Densovirinae
- Genus: Ambidensovirus
- Asteroid ambidensovirus 1
- Blattodean ambidensovirus 1
- Blattodean ambidensovirus 2
- Decapod ambidensovirus 1
- Dipteran ambidensovirus 1
- Hemipteran ambidensovirus 1
- Hemipteran ambidensovirus 2
- Hemipteran ambidensovirus 3
- Hymenopteran ambidensovirus 1
- Lepidopteran ambidensovirus 1
- Orthopteran ambidensovirus 1
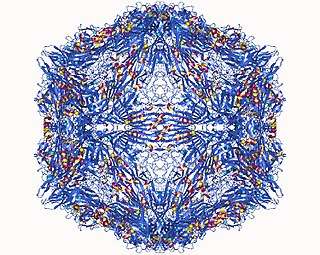
Structure
Virions consist of non-enveloped capsids that have a round appearance and display icosahedral symmetry.[7] The virions each have an isometric (and therefore spherical) nucleocapsid with a diameter of either 18–22 nm or 20–26 nm.[7] Sixty capsomers are present in each capsid.[7] The structure of each capsomer is described as "a quadrilateral 'kite-shaped' wedge"; the surface is said to have a rough appearance with small projections.[7][8] The center of capsids are sometimes visualized as appearing dark due to stain penetration in preparations where only a single species is retrieved. The virions do not appear to contain lipids. The buoyant density (in CsCl) of the virions is 1.4–1.44 g cm−3.[7]
| Genus | Structure | Symmetry | Capsid | Genomic arrangement | Genomic segmentation |
|---|---|---|---|---|---|
| Ambidensovirus | Icosahedral | T=1 | Non-enveloped | Linear | Segmented |
Genome
Ambidensoviruses have non-segmented genomes that contain a 5-6 kb linear ambisense single-stranded DNA and long inverted terminal repeats (550 bp). The ambisense genome encodes proteins on both the positive and negative sense strands,[5][9][10] Densoviruses use transcriptional regulation and post-transcriptional modification to produce different nonstructural proteins and structural proteins (VP).[11][8][10]
Life cycle
Viral replication is nuclear. Entry into the host cell is achieved by attachment to host receptors, which may be mediated by clathrin-mediated endocytosis[5] or clathrin-independent dynamin-dependent endocytosis.[12] Replication follows the rolling-hairpin model. DNA-templated transcription, with some alternative splicing mechanism is the method of transcription. Insects serve as the natural host.[5]
| Genus | Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|---|
| Ambidensovirus | Insects | Variable | Receptor-mediated endocytosis | uncertain | Nucleus | Nucleus | Unknown |
References
- "ICTV 10th Report (2018) Ambidensovirus".
- "ICTV 10th Report (2018) Densovirinae".
- "ICTV 10th Report (2018) Parvoviridae".
- Cotmore, SF; Agbandje-McKenna, M; Canuti, M; Chiorini, JA; Eis-Hubinger, A; Hughes, J; Mietzsch, M; Modha, S; Ogliastro, M; Pénzes, JJ; Pintel, DJ; Qiu, J; Soderlund-Venermo, M; Tattersall, P; Tijssen, P; and the ICTV Report Consortium (2019). "ICTV Virus Taxonomy Profile: Parvoviridae". Journal of General Virology. 100 (3): 367–368. doi:10.1099/jgv.0.001212. PMC 6537627. PMID 30672729.
- "Viral Zone". ExPASy. Retrieved 12 June 2015.
- ICTV. "Virus Taxonomy: 2017 Release". Retrieved 16 August 2018.
- "Densovirinae". ICTVdB—The Universal Virus Database, version 4. Retrieved 15 May 2007.
- Simpson AA, Chipman PR, Baker TS, Tijssen P, Rossmann MG (November 1998). "The structure of an insect parvovirus (Galleria mellonella densovirus) at 3.7 A resolution". Structure. 6 (11): 1355–67. doi:10.1016/S0969-2126(98)00136-1. PMC 4167665. PMID 9817847.
- Pham HT, Huynh OT, Jousset FX, Bergoin M, Tijssen P (August 2013). "Junonia coenia Densovirus (JcDNV) Genome Structure". Genome Announcements. 1 (4). doi:10.1128/genomeA.00591-13. PMC 3738899. PMID 23929483.
- Dumas B, Jourdan M, Pascaud AM, Bergoin M (November 1992). "Complete nucleotide sequence of the cloned infectious genome of Junonia coenia densovirus reveals an organization unique among parvoviruses". Virology. 191 (1): 202–22. doi:10.1016/0042-6822(92)90182-O. PMID 1413502.
- Bruemmer A, Scholari F, Lopez-Ferber M, Conway JF, Hewat EA (April 2005). "Structure of an insect parvovirus (Junonia coenia Densovirus) determined by cryo-electron microscopy". Journal of Molecular Biology. 347 (4): 791–801. doi:10.1016/j.jmb.2005.02.009. PMID 15769470.
- Wang Y, Gosselin Grenet AS, Castelli I, Cermenati G, Ravallec M, Fiandra L, Debaisieux S, Multeau C, Lautredou N, Dupressoir T, Li Y, Casartelli M, Ogliastro M (November 2013). "Densovirus crosses the insect midgut by transcytosis and disturbs the epithelial barrier function". Journal of Virology. 87 (22): 12380–91. doi:10.1128/JVI.01396-13. PMC 3807927. PMID 24027326.
Further reading
- Hewson I, Button JB, Gudenkauf BM, Miner B, Newton AL, Gaydos JK, et al. (December 2014). "Densovirus associated with sea-star wasting disease and mass mortality". Proceedings of the National Academy of Sciences of the United States of America. 111 (48): 17278–83. doi:10.1073/pnas.1416625111. PMC 4260605. PMID 25404293.