Mimiviridae
Mimiviridae is a family of viruses. Amoeba and other protists serve as natural hosts. The family is divided in up to 4 subfamilies.[1][2][3][4] Viruses in this family belong to the nucleocytoplasmic large DNA virus clade (NCLDV), also referred to as giant viruses.
| Mimiviridae | |
|---|---|
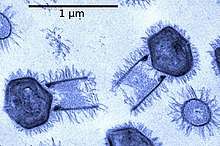 | |
| Tupanvirus | |
| Virus classification | |
| (unranked): | Virus |
| Phylum: | incertae sedis |
| Class: | incertae sedis |
| Order: | incertae sedis |
| Family: | Mimiviridae |
| Genera | |
| |
History
The first member of this family, Mimivirus, was discovered in 2003,[5] and the first complete genome sequence was published in 2004. [6] However, the mimivirus Cafeteria roenbergensis virus[7] was isolated and partially characterized in 1995[8], although the host was misidentified at the time, and the virus was designated BV-PW1.[7]
Taxonomy
Group: dsDNA
- Family: Mimiviridae
- *Cafeteria roenbergensis virus
- *Acanthamoeba polyphaga mimivirus
The genus is currently divided into three subfamilies.[2][3][9]
- One subfamily (Mimivirus, proposed names: Megavirinae or Megamimivirinae) is divided into three "lineages":
- A — Mimivirus group: includes Acanthamoeba polyphaga Mimivirus, Hirudovirus, Mamavirus, Kroon virus, Lentille virus, Terra2, Niemeyer virus, Samba virus.[10][11]
- B — Moumouvirus group: includes Moumouvirus, Saudi moumouvirus, Moumouvirus goulette, Monve virus (aka Moumouvirus monve), and Ochan virus.[12][10][13][11]
- C — Courdo11 virus group: includes Mont1,[10] Courdo7, Courdo11, Megavirus chilensis, LBA111, Powai lake megavirus and Terra1.[14][15]
- The majority of Mimiviridae appear to belong to this subfamily (Mimiviruses).[9]
- It is sometimes also referred to as Mimiviridae group I.[16]
- The second subfamily (Cafeteriavirus or Mimiviridae group II) includes the Cafeteria roenbergensis virus (CroV).[7]
- The Klosneuvirinae have been proposed as a third subfamily and are divided into four "lineages": Klosneuvirus, Indivirus, Catovirus and Hokovirus.[3] They seem to be closely related to the Mimivirus subfamily rather than the Cafeteriavirus subfamily (and so might be summarized in Mimivirus group I as well).[3]The first isolate from this group is Bodo saltans virus infecting the kinetoplastid Bodo saltans.[17]
- Tupanvirus strains have been discussed to comprise a sister group of mimiviruses.[4]
Furthermore, it has been proposed to extend Mimiviridae by an additional tentative group III (subfamily Mesomimivirinae aka legacy OLPG, Organic Lake Phycodna Group) that may consist of the following:
- Phaeocystis globosa virus (PgV, represented by PgV-16T strain) and Phaeocystis pouchetii virus (PpV, e. g. PpV 01)
- "Organic Lake Phycodnavirus" 1 and 2 (OLV1, OLV2, hosts of Organic Lake virophage)
- "Yellowstone Lake Mimivirus"[11][18] aka "Yellowstone Lake Phycodnavirus" 4 (YSLGV4)
- Chrysochromulina ericina virus (CeV, e. g. CeV 01)
- Aureococcus anophagefferens virus ([19]AaV)
- Pyramimonas orientalis virus (PoV)
- Tetraselmis virus (TetV-1)[20]
This group seems to be closely related to Mimiviridae rather than to Phycodnaviridae and therefore is sometimes referred to as a further subfamily candidate Mesomimivirinae. Sometimes the extended family Mimiviridae is referred to as Megaviridae although this has not been recognized by ICTV; alternatively the extended group may be referred to just as Mimiviridae.[3][21][22][23][24][16]
Although only a couple of members of this family have been described in detail it seems likely there are many more awaiting description and assignment[25][26] Unassigned members include Aureococcus anophagefferens virus (AaV), CpV-BQ2 and Terra2.
Structure
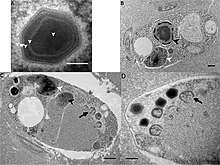
[17] Viruses in Mimiviridae have icosahedral and round geometries, with between T=972 and T=1141, or T=1200 symmetry. The diameter is around 400 nm, with a length of 125 nm. Genomes are linear and non-segmented, around 1200kb in length. The genome has 911 open reading frames.[1]
| Genus | Structure | Symmetry | Genomic arrangement | Genomic segmentation |
|---|---|---|---|---|
| Mimivirus | Icosahedral | T=972-1141 or T=1200 (H=19 +/- 1, K=19 +/- 1) | Linear | Monopartite |
| Klosneuvirus | ||||
| Cafeteriavirus | Icosahedral | T=499 | Linear | Monopartite |
| Tupanvirus | Tailed |
Life cycle
Replication follows the DNA strand displacement model. DNA-templated transcription is the method of transcription. Amoeba serve as the natural host.[1]
| Genus | Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|---|
| Mimivirus | Amoeba | None | Unknown | Unknown | Unknown | Unknown | Passive diffusion |
| Klosneuvirus | microzooplankton | None | Unknown | Unknown | Unknown | Cytoplasm | Passive diffusion |
| Cafeteriavirus | microzooplankton | None | Unknown | Unknown | Unknown | Cytoplasm | Passive diffusion |
Molecular biology
Within the genome of Lentille virus integrated genome of a virophage (Sputnik 2) and a transpoviron—a mobile genetic element—have been reported. Transpovirons are linear DNA elements of about 7 kilobases that encompass six to eight protein coding genes, two of which are homologous to virophage genes. Broad spectrum of mimiviridae virophage allows its isolation using a mimivirus reporter.[14]
Clinical
Mimiviruses have been associated with pneumonia but their significance if any is currently unknown.[27] The only virus of this family isolated from a human to date is LBA 111.[28] Mimivirus has also been implicated in rheumatoid arthritis.[29]
References
- "Viral Zone". ExPASy. Retrieved 15 June 2015.
- ICTV. "Virus Taxonomy: 2014 Release". Retrieved 15 June 2015.
- Schulz, Frederik; Yutin, Natalya; Ivanova, Natalia N.; Ortega, Davi R.; Lee, Tae Kwon; Vierheilig, Julia; Daims, Holger; Horn, Matthias; Wagner, Michael (7 April 2017). "Giant viruses with an expanded complement of translation system components" (PDF). Science. 356 (6333): 82–85. Bibcode:2017Sci...356...82S. doi:10.1126/science.aal4657. ISSN 0036-8075. PMID 28386012., UCPMS ID: 1889607, PDF
- Abrahão, Jônatas; Silva, Lorena; Silva, Ludmila Santos; Khalil, Jacques Yaacoub Bou; Rodrigues, Rodrigo; Arantes, Thalita; Assis, Felipe; Boratto, Paulo; Andrade, Miguel; Kroon, Erna Geessien; Ribeiro, Bergmann; Bergier, Ivan; Seligmann, Herve; Ghigo, Eric; Colson, Philippe; Levasseur, Anthony; Kroemer, Guido; Raoult, Didier; Scola, Bernard La (27 February 2018). "Tailed giant Tupanvirus possesses the most complete translational apparatus of the known virosphere". Nature Communications. 9 (1): 749. Bibcode:2018NatCo...9..749A. doi:10.1038/s41467-018-03168-1. PMC 5829246. PMID 29487281. Fig. 4 and §Discussion: "Considering that tupanviruses comprise a sister group to amoebal mimiviruses…"
- Suzan-Monti, M; La Scola, B; Raoult, D (2006). "Genomic and evolutionary aspects of Mimivirus". Virus Res. 117 (1): 145–155. doi:10.1016/j.virusres.2005.07.011. PMID 16181700.
- Raoult, D.; Audic, S; Robert, C; Abergel, C; Renesto, P; Ogata, H; La Scola, B; Suzan, M; Claverie, JM (2004). "The 1.2-Megabase Genome Sequence of Mimivirus". Science. 306 (5700): 1344–50. doi:10.1126/science.1101485. PMID 15486256.
- Matthias G. Fischer; Michael J. Allen; William H. Wilson; Curtis A. Suttle (2010). "Giant virus with a remarkable complement of genes infects marine zooplankton". Proceedings of the National Academy of Sciences. 107 (45): 19508–13. Bibcode:2010PNAS..10719508F. doi:10.1073/pnas.1007615107. PMC 2984142. PMID 20974979.
- D.R. Garza; C.A. Suttle (1995). "Large double-stranded DNA viruses which cause the lysis of a marine heterotrophic nanoflagellate (Bodo sp.) occur in natural marine viral communities". Aquatic Microbial Ecology. 9 (3): 203–210. doi:10.3354/ame009203.
- Colson P, Fournous G, Diene SM, Raoult D (2013). "Codon usage, amino acid usage, transfer RNA and amino-acyl-tRNA synthetases in Mimiviruses". Intervirology. 56 (6): 364–75. doi:10.1159/000354557. PMID 24157883.
- Gaia M, Benamar S, Boughalmi M, Pagnier I, Croce O, Colson P, Raoult D, La Scola B (2014). "Zamilon, a novel virophage with Mimiviridae host specificity". PLoS ONE. 9 (4): e94923. doi:10.1371/journal.pone.0094923. PMC 3991649. PMID 24747414.
- See also Abrahão & et al. 2018, fig. 4 on p. 5
- Desnues C, La Scola B, Yutin N, Fournous G, Robert C, Azza S, Jardot P, Monteil S, Campocasso A, Koonin EV, Raoult D (October 2012). "Provirophages and transpovirons as the diverse mobilome of giant viruses". Proc. Natl. Acad. Sci. U.S.A. 109 (44): 18078–83. doi:10.1073/pnas.1208835109. PMC 3497776. PMID 23071316.
- Yutin N, Wolf YI, Koonin EV (October 2014). "Origin of giant viruses from smaller DNA viruses not from a fourth domain of cellular life". Virology. 466-467: 38–52. doi:10.1016/j.virol.2014.06.032. PMC 4325995. PMID 25042053.
- Gaia M, Pagnier I, Campocasso A, Fournous G, Raoult D, La Scola B (2013). "Broad spectrum of mimiviridae virophage allows its isolation using a mimivirus reporter". PLoS ONE. 8 (4): e61912. doi:10.1371/journal.pone.0061912. PMC 3626643. PMID 23596530.
- For LBA111 and Powai lake megavirus see also Abrahão & et al. 2018, fig. 4 on p. 5
- Zhang W, Zhou J, Liu T, Yu Y, Pan Y, Yan S, Wang Y (October 2015). "Four novel algal virus genomes discovered from Yellowstone Lake metagenomes". Sci Rep. 5: 15131. doi:10.1038/srep15131. PMC 4602308. PMID 26459929.
- Deeg, C.M.; Chow, E.C.T.; Suttle, C.A. (2018). "The kinetoplastid-infecting Bodo saltans virus (BsV), a window into the most abundant giant viruses in the sea". eLife. 7: e33014. doi:10.7554/eLife.33014. PMC 5871332. PMID 29582753.
- NCBI Complete genomes: Viruses, look for 'Yellowstone Lake'
- Moniruzzaman, Mohammad; LeCleir, Gary R.; Brown, Christopher M.; Gobler, Christopher J.; Bidle, Kay D.; Wilson, William H.; Wilhelm, Steven W. (2014). "Genome of brown tide virus (AaV), the little giant of the Megaviridae, elucidates NCLDV genome expansion and host–virus coevolution". Virology. 466-467: 60–70. doi:10.1016/j.virol.2014.06.031. PMID 25035289.
- Schvarcz CR, Steward GF (May 2018). "A giant virus infecting green algae encodes key fermentation genes". Virology. 518: 423–433. doi:10.1016/j.virol.2018.03.010. PMID 29649682. Lay summary – ScienceDaily.
- Koonin EV, Krupovic M, Yutin N (April 2015). "Evolution of double-stranded DNA viruses of eukaryotes: from bacteriophages to transposons to giant viruses". Ann. N. Y. Acad. Sci. 1341: 10–24, see Figure 3. doi:10.1111/nyas.12728. PMC 4405056. PMID 25727355.
- Yutin N, Colson P, Raoult D, Koonin EV (April 2013). "Mimiviridae: clusters of orthologous genes, reconstruction of gene repertoire evolution and proposed expansion of the giant virus family". Virol. J. 10: 106. doi:10.1186/1743-422X-10-106. PMC 3620924. PMID 23557328.
- Blog of Carolina Reyes, Kenneth Stedman: Are Phaeocystis globosa viruses (OLPG) and Organic Lake phycodnavirus a part of the Phycodnaviridae or Mimiviridae?, on ResearchGate, Jan. 8, 2016
- Maruyama F, Ueki S (2016). "Evolution and Phylogeny of Large DNA Viruses, Mimiviridae and Phycodnaviridae Including Newly Characterized Heterosigma akashiwo Virus". Front Microbiol. 7: 1942. doi:10.3389/fmicb.2016.01942. PMC 5127864. PMID 27965659.
- Ghedin E, Claverie JM (August 2005). "Mimivirus relatives in the Sargasso sea". Virol. J. 2: 62. doi:10.1186/1743-422X-2-62. PMC 1215527. PMID 16105173.
- Monier A, Claverie JM, Ogata H (2008). "Taxonomic distribution of large DNA viruses in the sea". Genome Biol. 9 (7): R106. doi:10.1186/gb-2008-9-7-r106. PMC 2530865. PMID 18598358.
- Saadi H, Pagnier I, Colson P, Cherif JK, Beji M, Boughalmi M, Azza S, Armstrong N, Robert C, Fournous G, La Scola B, Raoult D (August 2013). "First isolation of Mimivirus in a patient with pneumonia". Clin. Infect. Dis. 57 (4): e127–34. doi:10.1093/cid/cit354. PMID 23709652.
- Yoosuf N, Pagnier I, Fournous G, Robert C, La Scola B, Raoult D, Colson P (April 2014). "Complete genome sequence of Courdo11 virus, a member of the family Mimiviridae". Virus Genes. 48 (2): 218–23. doi:10.1007/s11262-013-1016-x. PMID 24293219.
- Shah, N.; Hulsmeier, A. J.; Hochhold, N.; Neidhart, M.; Gay, S.; Hennet, T. (2013). "Exposure to Mimivirus Collagen Promotes Arthritis". Journal of Virology. 88 (2): 838–45. doi:10.1128/JVI.03141-13. PMC 3911627. PMID 24173233.