Staphylococcus epidermidis
Staphylococcus epidermidis is a Gram-positive bacterium, and one of over 40 species belonging to the genus Staphylococcus.[1] It is part of the normal human flora, typically the skin flora, and less commonly the mucosal flora.[2] It is a facultative anaerobic bacteria. Although S. epidermidis is not usually pathogenic, patients with compromised immune systems are at risk of developing infection. These infections are generally hospital-acquired.[3] S. epidermidis is a particular concern for people with catheters or other surgical implants because it is known to form biofilms that grow on these devices.[4] Being part of the normal skin flora, S. epidermidis is a frequent contaminant of specimens sent to the diagnostic laboratory.[5]

| Staphylococcus epidermidis | |
|---|---|
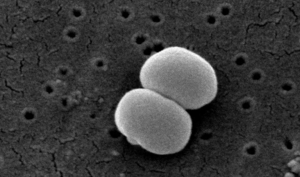 | |
| Scanning electron image of S. epidermidis. | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | |
| Class: | |
| Order: | |
| Family: | |
| Genus: | |
| Species: | S. epidermidis |
| Binomial name | |
| Staphylococcus epidermidis (Winslow & Winslow 1908) Evans 1916 | |
| Synonyms | |
|
Staphylococcus albus Rosenbach 1884 | |
Etymology
'Staphylococcus' - bunch of grape-like berries, 'epidermidis' - of the epidermis.[6]
Discovery
Friedrich Julius Rosenbach distinguished S. epidermidis from S. aureus in 1884, initially naming S. epidermidis as S. albus.[7] He chose aureus and albus since the bacteria formed yellow and white colonies, respectively.
Cellular morphology and biochemistry
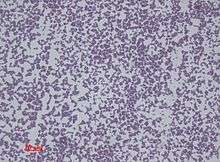
S. epidermidis is a very hardy microorganism, consisting of nonmotile, Gram-positive cocci, arranged in grape-like clusters. It forms white, raised, cohesive colonies about 1–2 mm in diameter after overnight incubation, and is not hemolytic on blood agar.[4] It is a catalase-positive,[8] coagulase-negative, facultative anaerobe that can grow by aerobic respiration or by fermentation. Some strains may not ferment.[9]
Biochemical tests indicate this microorganism also carries out a weakly positive reaction to the nitrate reductase test. It is positive for urease production, is oxidase negative, and can use glucose, sucrose, and lactose to form acid products. In the presence of lactose, it will also produce gas. S. epidermidis does not possess the gelatinase enzyme, so it cannot hydrolyze gelatin. It is sensitive to novobiocin, providing an important test to distinguish it from Staphylococcus saprophyticus, which is coagulase-negative, as well, but novobiocin-resistant.[3]
Similar to those of S. aureus, the cell walls of S. epidermidis have a transferrin-binding protein that helps the organism obtain iron from transferrin. The tetramers of a surface exposed protein, glyceraldehyde-3-phosphate dehydrogenase, are believed to bind to transferrin and remove its iron. Subsequent steps include iron being transferred to surface lipoproteins, then to transport proteins which carry the iron into the cell.[4]
Virulence and antibiotic resistance
The ability to form biofilms on plastic devices is a major virulence factor for S. epidermidis. One probable cause is surface proteins that bind blood and extracellular matrix proteins. It produces an extracellular material known as polysaccharide intercellular adhesin (PIA), which is made up of sulfated polysaccharides. It allows other bacteria to bind to the already existing biofilm, creating a multilayer biofilm. Such biofilms decrease the metabolic activity of bacteria within them. This decreased metabolism, in combination with impaired diffusion of antibiotics, makes it difficult for antibiotics to effectively clear this type of infection.[4] S. epidermidis strains are often resistant to antibiotics, including rifamycin, fluoroquinolones, gentamicin, tetracycline, clindamycin, and sulfonamides.[10] Methicillin resistance is particularly widespread, with 75-90% of hospital isolates resistance to methicillin.[10] Resistant organisms are most commonly found in the intestine, but organisms living freely on the skin can also become resistant due to routine exposure to antibiotics secreted in sweat.
Disease
As mentioned above, S. epidermidis causes biofilms to grow on plastic devices placed within the body.[11] This occurs most commonly on intravenous catheters and on medical prostheses.[12] Infection can also occur in dialysis patients or anyone with an implanted plastic device that may have been contaminated. It also causes endocarditis, most often in patients with defective heart valves. In some other cases, sepsis can occur in hospital patients.
Antibiotics are largely ineffective in clearing biofilms. The most common treatment for these infections is to remove or replace the infected implant, though in all cases, prevention is ideal. The drug of choice is often vancomycin, to which rifampin or an aminoglycoside can be added. Hand washing has been shown to reduce the spread of infection.
Preliminary research also indicates S. epidermidis is universally found inside affected acne vulgaris pores, where Cutibacterium acnes is normally the sole resident.[13]
Identification
The normal practice of detecting S. epidermidis is by using appearance of colonies on selective media, bacterial morphology by light microscopy, catalase and slide coagulase testing. On the Baird-Parker agar with egg yolk supplement, colonies appear small and black. Increasingly, techniques such as quantitative PCR are being employed for the rapid detection and identification of Staphylococcus strains.[14][15] Normally, sensitivity to desferrioxamine can also be used to distinguish it from most other staphylococci, except in the case of Staphylococcus hominis, which is also sensitive.[16] In this case, the production of acid from trehalose by S. hominis can be used to tell the two species apart.
See also
- Biofilms
- Microbiology
- Staphylococcus
Notes and references
- Schleifer, K. H.; Kloos, W. E. (1975). "Isolation and Characterization of Staphylococci from Human Skin I. Amended Descriptions of Staphylococcus epidermidis and Staphylococcus saprophyticus and Descriptions of Three New Species Staphylococcus cohnii, Staphylococcus haemolyticus, and Staphylococcus xylosus". International Journal of Systematic Bacteriology. 25 (1): 50–61. doi:10.1099/00207713-25-1-50. ISSN 0020-7713.
- Fey, P. D.; Olson, M. E. (2010). "Current concepts in biofilm formation of Staphylococcus epidermidis". Future Microbiology. 5 (6): 917–933. doi:10.2217/fmb.10.56. PMC 2903046. PMID 20521936.
- Levinson, W. (2010). Review of Medical Microbiology and Immunology (11th ed.). pp. 94–99.
- Salyers, Abigail A. & Whitt, Dixie D. (2002). Bacterial Pathogenesis: A Molecular Approach, 2nd ed. Washington, D.C.: ASM Press. ISBN 978-1-55581-171-6.
- Queck SY & Otto M (2008). "Staphylococcus epidermidis and other Coagulase-Negative Staphylococci". Staphylococcus: Molecular Genetics. Caister Academic Press. ISBN 978-1-904455-29-5.
- "VetBact".
- Friedrich Julius Rosenbach at Who Named It?
- "Todar's Online Textbook of Bacteriology: Staphylococcus aureus and Staphylococcal Disease". Kenneth Todar, PhD. Retrieved Dec 7, 2013.
- "Bacteria Genomes - STAPHYLOCOCCUS EPIDERMIDIS". Karyn's Genomes. EMBL-EBI. Retrieved December 23, 2011.
- Otto M (August 2010). "Staphylococcus epidermidis - the "accidental" pathogen". Nature Reviews Microbiology. 7 (8): 555–567. doi:10.1038/nrmicro2182. PMC 2807625. PMID 19609257.
- Otto M (2009), "Staphylococcus epidermidis — the 'accidental' pathogen", Nature Reviews Microbiology, 7 (8): 555–567, doi:10.1038/nrmicro2182, PMC 2807625, PMID 19609257
- Hedin G (1993), "Staphylococcus epidermidis — hospital epidemiology and the detection of methicillin resistance", Scandinavian Journal of Infectious Diseases Supplementum, 90: 1–59, PMID 8303217
- Bek-Thomson, M.; et al. (2008). "Acne is Not Associated with Yet-Uncultured Bacteria". Journal of Clinical Microbiology. 46 (10): 3355–3360. doi:10.1128/JCM.00799-08. PMC 2566126. PMID 18716234.
- Francois P & Schrenzel J (2008). "Rapid Diagnosis and Typing of Staphylococcus aureus". Staphylococcus: Molecular Genetics. Caister Academic Press. ISBN 978-1-904455-29-5.
- Mackay IM (editor). (2007). Real-Time PCR in Microbiology: From Diagnosis to Characterization. Caister Academic Press. ISBN 978-1-904455-18-9.
- Antunes, Ana Lúcia Souza; Secchi, Carina; Reiter, Keli Cristine; Perez, Leandro Reus Rodrigues; Freitas, Ana Lúcia Peixoto De; D'azevedo, Pedro Alves (2008-01-01). "Feasible identification of Staphylococcus epidermidis using desferrioxamine and fosfomycin disks". APMIS. 116 (1): 16–20. doi:10.1111/j.1600-0463.2008.00796.x. ISSN 1600-0463. PMID 18254775.
External links
- Type strain of Staphylococcus epidermidis at BacDive - the Bacterial Diversity Metadatabase
- Teruaki Nakatsuji et al.: A commensal strain of Staphylococcus epidermidis protects against skin neoplasia, in: Science Advances; 28th of Feb., 2018; Vol. 4, No. 2, DOI:10.1126/sciadv.aao4502
- Barros J, Grenho L, Manuel C, Ferreira C, Melo L, Nunes O, Monteiro F, Ferraz M. Influence of Nanohydroxyapatite Surface Properties on Staphylococcus epidermidis Biofilm Formation. Journal of Biomaterials Applications. 2013;28(9):1325–1335.
- Dong Y, Glaser K, Schlegel N, Claus H, Speer CP. An underestimated pathogen: Staphylococcus epidermidis induces pro-inflammatory responses in human alveolar epithelial cells. Cytokine. 2019;123:154761.
- Feng G, Cheng Y, Worobo R, Borca‐Tasciuc D, Moraru C. Nanoporous anodic alumina reduces Staphylococcus biofilm formation. Letters in Applied Microbiology. 2019;69(4):246–251.
- Gill SR, Fouts DE, Archer GL, Mongodin EF, Deboy RT, Ravel J, Paulsen IT, Kolonay JF, Brinkac L, Beanan M, et al. Insights on Evolution of Virulence and Resistance from the Complete Genome Analysis of an Early Methicillin-Resistant Staphylococcus aureus Strain and a Biofilm-Producing Methicillin-Resistant Staphylococcus epidermidis Strain. Journal of Bacteriology. 2005;187(7):2426–2438.
- Gotz F. Staphylococcus and biofilms. Molecular Microbiology. 2002 [accessed 2019 Oct 2]; 43(6):1367–1378. doi:10.1046/j.1365-2958.2002.02827.x
- Haidamak J, Santos GDD, Bruna Jacomel Favoreto De Souza Lima, Soares VM, Menezes RVD, Bisson AA, Talevi AS, Gomes RR, Vicente VA, Valero MA, et al. Scalp microbiota alterations in children with pediculosis. Infection, Genetics and Evolution. 2019;73:322–331.
- Izano EA, Amarante MA, Kher WB, Kaplan JB. Differential Roles of Poly-N-Acetylglucosamine Surface Polysaccharide and Extracellular DNA in Staphylococcus aureus and Staphylococcus epidermidis Biofilms. Applied and Environmental Microbiology. 2007;74(2):470–476.
- Méric G, Miragaia M, Been MD, Yahara K, Pascoe B, Mageiros L, Mikhail J, Harris LG, Wilkinson TS, Rolo J, et al. Ecological Overlap and Horizontal Gene Transfer in Staphylococcus aureus and Staphylococcus epidermidis. Genome Biology and Evolution. 2015;7(5):1313–1328.
- Otto M. Staphylococcus epidermidis — the accidental pathogen. Nature Reviews Microbiology. 2009 [accessed 2019 Oct 2];7(8):555–567. doi:10.1038/nrmicro2182
- Qin Z, Ou Y, Yang L, Zhu Y, Tolker-Nielsen T, Molin S, Qu D. Role of autolysin-mediated DNA release in biofilm formation of Staphylococcus epidermidis. Microbiology. 2007 [accessed 2019 Oct 2];153(7):2083–2092. doi:10.1099/mic.0.2007/006031-0
- Schaeffer CR, Hoang T-MN, Sudbeck CM, Alawi M, Tolo IE, Robinson DA, Horswill AR, Rohde H, Fey PD. Versatility of Biofilm Matrix Molecules in Staphylococcus epidermidis Clinical Isolates and Importance of Polysaccharide Intercellular Adhesin Expression during High Shear Stress. mSphere. 2016;1(5).
- Shahrooei M, Hira V, Khodaparast L, Khodaparast L, Stijlemans B, Kucharíková S, Burghout P, Hermans PWM, Eldere JV. Vaccination with SesC Decreases Staphylococcus epidermidis Biofilm Formation. Infection and Immunity. 2012;80(10):3660–3668.