Sporulation in Bacillus subtilis
Bacillus subtilis is a rod-shaped, Gram-positive bacteria that is naturally found in soil and vegetation, and is known for its ability to form a small, tough, protective and metabolically dormant endospore. B. subtilis can divide symmetrically to make two daughter cells (binary fission), or asymmetrically, producing a single endospore that is resistant to environmental factors such as heat, desiccation, radiation and chemical insult which can persist in the environment for long periods of time. The endospore is formed at times of nutritional stress, allowing the organism to persist in the environment until conditions become favourable. The process of endospore formation has profound morphological and physiological consequences: radical post-replicative remodelling of two progeny cells, accompanied eventually by cessation of metabolic activity in one daughter cell (the spore) and death by lysis of the other (the ‘mother cell’).
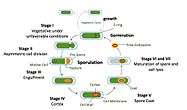
Overview
Commitment to sporulation
Although sporulation in B. subtilis is induced by starvation, the sporulation developmental program is not initiated immediately when growth slows due to nutrient limitation. A variety of alternative responses can occur, including the activation of flagellar motility to seek new food sources by chemotaxis, the production of antibiotics to destroy competing soil microbes, the secretion of hydrolytic enzymes to scavenge extracellular proteins and polysaccharides, or the induction of ‘competence’ for uptake of exogenous DNA for consumption, with the occasional side-effect that new genetic information is stably integrated. Sporulation is the last-ditch response to starvation and is suppressed until alternative responses prove inadequate. Even then, certain conditions must be met such as chromosome integrity, the state of chromosomal replication, and the functioning of the Krebs cycle.[1]
Nature of regulation
Sporulation requires a great deal of time and energy and is essentially irreversible,[2] making it crucial for a cell to monitor its surroundings efficiently and ensure that sporulation is embarked upon at only the most appropriate times. The wrong decision can be catastrophic: a vegetative cell will die if the conditions are too harsh, while bacteria forming spores in an environment which is conducive to vegetative growth will be out competed.[3] In short, initiation of sporulation is a very tightly regulated network with numerous checkpoints for efficient control.
Control points
Two transcriptional regulators, σH and Spo0A, play key roles in initiation of sporulation. Several additional proteins participate, mainly by controlling the accumulated concentration of Spo0A~P. Spo0A lies at the end of a series of inter-protein phosphotransfer reactions, Kin–Spo0F–Spo0B–Spo0A, termed as a ‘phosphorelay’. The regulation of these various factors controlling the accumulated concentration of Spo0A~P, and their interactions are described in detail in Figure2:
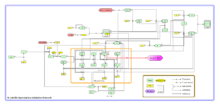
In the table below, the term 'Activators' refers to genes/proteins that ultimately result in initiation of sporulation and 'Repressors' refers to the ones that inhibit this initiation of sporulation.
| Activators | Regulation | Repressors | Regulation |
|---|---|---|---|
| KinA | Transfers phosphate to the Spo0F | Sda | Blocks autophosphorylation of KinA |
| KinB | Transfers phosphate to the Spo0F | KipI | Blocks autophosphorylation of KinA |
| Spo0A | Activates several key sporulation-specific genes | Spo0A | Negatively controls transcription of abrB |
| Spo0H (σH) | Activates phrE gene | ComA | Activates competence |
| Spo0B | Phosphotransferase initiation | SinR | Negative regulation of kinB |
| Spo0F | Phosphotransferase initiation | RapA, RapB, RapE, and RapH | Dephosphorylation of Spo0F~P |
| ComA | Activates phrA | Spo0E, YisI, and YnzD | Dephosphorylation of Spo0A~P |
| SinI | Antagonist of SinR | GTP-bound CodY | Inhibits rapA-phrA |
| KipA | Inhibits kipI gene | AbrB | Encodes a repressor of spo0H |
| PhrA | Suppresses dephosphorylation activity of RapA | Hpr | Overexpression inhibits sporulation |
| PhrE | Inhibits RapE | DnaA | Over expression of Sda |
| PhrH | Inhibits RapH |
Homologous networks in eukaryotes
Spores form a part of the life cycles of a diverse range of organisms such as many bacteria, plants, algae, fungi and some protozoa. In Saccharomyces cerevisiae (Kingdom Fungi), the set of early genes activating sporulation is induced by Ime1 (Inducer of Meiosis 1) and a regulator of middle genes is Ndt80p.[4] Similarly, in Dictyostelium discoideum (slime mold), SDF-2 and cytokinins are secreted during late development to trigger signal transduction pathways that lead to an increase in the activity of the cAMP-dependent protein kinase, PKA, which triggers rapid encapsulation as well as ensuring spore dormancy.[5]
See also
- Bacillus
- Sporophyte
- Bacillus subtilis
References
- Stephens, Craig (1998). "Bacterial sporulation: A question of commitment?". Current Biology. 8 (2): R45–8. doi:10.1016/S0960-9822(98)70031-4. PMID 9427639.
- Piggot, PJ; Coote, JG (1976). "Genetic aspects of bacterial endospore formation". Bacteriological Reviews. 40 (4): 908–62. PMC 413989. PMID 12736.
- Jabbari, Sara; Heap, John T.; King, John R. (2010). "Mathematical Modelling of the Sporulation-Initiation Network in Bacillus Subtilis Revealing the Dual Role of the Putative Quorum-Sensing Signal Molecule PhrA" (PDF). Bulletin of Mathematical Biology. 73 (1): 181–211. doi:10.1007/s11538-010-9530-7. PMID 20238180.
- Piekarska, Iga; Rytka, Joanna; Rempola, Bozenna (2010). "Regulation of sporulation in the yeast Saccharomyces cerevisiae" (PDF). Acta Biochimica Polonica. 57 (3): 241–50. PMID 20842291.
- Anjard, C.; Loomis, W. F. (2008). "Cytokinins induce sporulation in Dictyostelium". Development. 135 (5): 819–27. doi:10.1242/dev.018051. PMID 18216168.