Methylotroph
Methylotrophs are a diverse group of microorganisms that can use reduced one-carbon compounds, such as methanol or methane, as the carbon source for their growth; and multi-carbon compounds that contain no carbon-carbon bonds, such as dimethyl ether and dimethylamine. This group of microorganisms also includes those capable of assimilating reduced one-carbon compounds by way of carbon dioxide using the ribulose bisphosphate pathway.[1] These organisms should not be confused with methanogens which on the contrary produce methane as a by-product from various one-carbon compounds such as carbon dioxide. Some methylotrophs can degrade the greenhouse gas methane, and in this case they are called methanotrophs. The abundance, purity, and low price of methanol compared to commonly used sugars make methylotrophs competent organisms for production of amino acids, vitamins, recombinant proteins, single-cell proteins, co-enzymes and cytochromes.
Metabolism
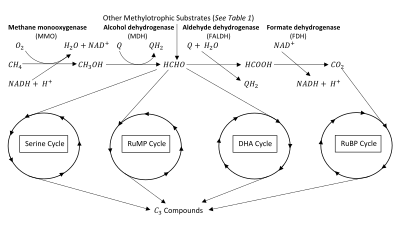
The key intermediate in methylotrophic metabolism is formaldehyde, which can be diverted to either assimilatory or dissimilatory pathways.[2] Methylotrophs produce formaldehyde through oxidation of methanol and/or methane. Methane oxidation requires the enzyme methane monooxygenase (MMO).[3][4] Methylotrophs with this enzyme are given the name methanotrophs. The oxidation of methane (or methanol) can be assimilatory or dissimilatory in nature (see figure). If dissimilatory, the formaldehyde intermediate is oxidized completely into to produce reductant and energy.[5][6] If assimilatory, the formaldehyde intermediate is used to synthesize a 3-Carbon () compound for the production of biomass.[2][7] Many methylotrophs use multi-carbon compounds for anabolism, thus limiting their use of formaldehyde to dissimilatory processes, however methanotrophs are generally limited to only metabolism.[2][5]
| Single Carbon Compounds | Chemical Formula | Multi-Carbon Compounds | Chemical Formula |
|---|---|---|---|
| Carbon monoxide | Dimethyl ether | ||
| Formaldehyde | Dimethylamine | ||
| Formamide | Dimethyl sulfide | ||
| Formic acid | Tetramethylammonium | ||
| Methane | Trimethylamine | ||
| Methanol | Trimethylamine N-oxide | ||
| Methylamine | Trimethylsuphonium | ||
| Methyl halide |
Catabolism
Methylotrophs use the electron transport chain to conserve energy produced from the oxidation of compounds. An additional activation step is required in methanotrophic metabolism to allow degradation of chemically-stable methane. This oxidation to methanol is catalyzed by MMO, which incorporates one oxygen atom from into methane and reduces the other oxygen atom to water, requiring two equivalents of reducing power.[4][5] Methanol is then oxidized to formaldehyde through the action of methanol dehydrogenase (MDH) in bacteria,[12] or a non-specific alcohol oxidase in yeast.[13] Electrons from methanol oxidation are passed to a membrane-associated quinone of the electron transport chain to produce .[14]
In dissimilatory processes, formaldehyde is completely oxidized to and excreted. Formaldehyde is oxidized to formate via the action of Formaldehyde dehydrogenase (FALDH), which provides electrons directly to a membrane associated quinone of the electron transport chain, usually cytochrome b or c.[2][5] In the case of associated dehydrogenases, is produced.[7]
Finally, formate is oxidized to by cytoplasmic or membrane-bound Formate dehydrogenase (FDH), producing [15] and .
Anabolism
The main metabolic challenge for methylotrophs is the assimilation of single carbon units into biomass. Through de novo synthesis, methylotrophs must form carbon-carbon bonds between 1-Carbon () molecules. This is an energy intensive process, which facultative methylotrophs avoid by using a range of larger organic compounds.[16] However, obligate methylotrophs must assimilate molecules.[2][5] There are four distinct assimilation pathways with the common theme of generating one molecule.[2] Bacteria use three of these pathways[7][11] while Fungi use one.[17] All four pathways incorporate 3 molecules into multi-carbon intermediates, then cleave one intermediate into a new molecule. The remaining intermediates are rearranged to regenerate the original multi-carbon intermediates.
Bacteria
Each species of methylotrophic bacteria has a single dominant assimilation pathway.[5] The three characterized pathways for carbon assimilation are the ribulose monophosphate (RuMP) and serine pathways of formaldehyde assimilation as well as the ribulose bisphosphate (RuBP) pathway of CO2 assimilation.[2][7][11][18]
Ribulose bisphosphate (RuBP) cycle
Unlike the other assimilatory pathways, bacteria using the RuBP pathway derive all of their organic carbon from assimilation.[5][19] This pathway was first elucidated in photosynthetic autotrophs and is better known as the Calvin Cycle.[19][20] Shortly thereafter, methylotrophic bacteria who could grow on reduced compounds were found using this pathway.[21]
First, 3 molecules of ribulose 5-phosphate are phosphorylated to ribulose 1,5-bisphosphate (RuBP). The enzyme ribulose bisphosphate carboxylase (RuBisCO) carboxylates these RuBP molecules which produces 6 molecules of 3-phosphoglycerate (PGA). The enzyme phosphoglycerate kinase phosphorylates PGA into 1,3-diphosphoglycerate (DPGA). Reduction of 6 DPGA by the enzyme glyceraldehyde phosphate dehydrogenase generates 6 molecules of the compound glyceraldehyde-3-phosphate (GAP). One GAP molecule is diverted towards biomass while the other 5 molecules regenerate the 3 molecules of ribulose 5-phosphate.[7][20]
Ribulose monophosphate (RuMP) cycle
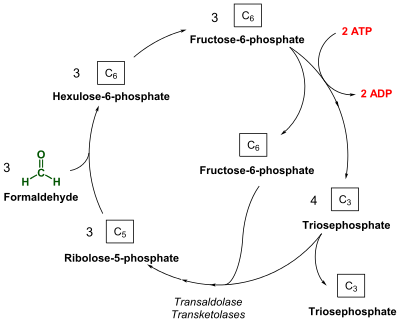
A new pathway was suspected when RuBisCO was not found in the methanotroph Methylmonas methanica.[22] Through radio-labelling experiments, it was shown that M. methanica used the Ribulose monophate (RuMP) pathway. This has led researchers to propose that the RuMP cycle may have preceded the RuBP cycle.[5]
Like the RuBP cycle, this cycle begins with 3 molecules of ribulose-5-phosphate. However, instead of phosphorylating ribulose-5-phosphate, 3 molecules of formaldehyde form a C-C bond through an aldol condensation, producing 3 molecules of 3-hexulose 6-phosphate (hexulose phosphate). One of these molecules of hexulose phosphate is converted into GAP and either pyruvate or dihydroxyacetone phosphate (DHAP). The pyruvate or DHAP is used towards biomass while the other 2 hexulose phosphate molecules and the molecule of GAP are used to regenerate the 3 molecules of ribulose-5-phosphate.[6][22]
Serine cycle
Unlike the other assimilatory pathways, the serine cycle uses carboxylic acids and amino acids as intermediates instead of carbohydrates.[5][23] First, 2 molecules of formaldehyde are added to 2 molecules of the amino acid glycine. This produces two molecules of the amino acid serine, the key intermediate of this pathway. These serine molecules eventually produce 2 molecules of 2-phosphoglycerate, with one molecule going towards biomass and the other being used to regenerate glycine. Notably, the regeneration of glycine requires a molecule of as well, therefore the Serine pathway also differs from the other 3 pathways by its requirement of both formaldehyde and .[22][23]
Yeasts
Methylotrophic yeast metabolism differs from bacteria primarily on the basis of the enzymes used and the carbon assimilation pathway. Unlike bacteria which use bacterial MDH, methylotrophic yeasts oxidize methanol in their peroxisomes with a non-specific alcohol oxidase. This produces formaldehyde as well as hydrogen peroxide.[24][25] Compartmentalization of this reaction in peroxisomes likely sequesters the hydrogen peroxide produced. Catalase is produced in the peroxisomes to deal with this harmful by-product.[17][24]
Dihydroxyacteone (DHA) cycle
The dihydroxyacetone (DHA) pathway, also known as the xylulose monophosphate (XuMP) pathway, is found exclusively in yeast.[24][26] This pathway assimilates three molecules of formaldehyde into 1 molecule of DHAP using 3 molecules of xylulose 5-phosphate as the key intermediate.
DHA synthase acts as a transferase (transketolase) to transfer part of xylulose 5-phosphate to DHA. Then these 3 molecules of DHA are phosphorylated to DHAP by triokinase. Like the other cycles, 3 molecules are produced with 1 molecule being directed for use as cell material. The other 2 molecules are used to regenerate xylulose 5-phosphate.[27]
Environmental Implications
As key players in the carbon cycle, methylotrophs work to reduce global warming primarily through the uptake of methane and other greenhouse gases. In aqueous environments, methanogenic bacteria produce 40-50% of the world's methane. Symbiosis between methanogens and methanotrophic bacteria greatly decreases the amount of methane released into the atmosphere.[28]
The use of methylotrophs in the agricultural sector is another way in which they can potentially impact the environment. Traditional chemical fertilizers supply nutrients not readily available from soil but can have some negative environmental impacts and are costly to produce.[29] Methylotrophs have high potential as alternative biofertilizers and bioinoculants due to their ability form mutualistic relationships with several plant species.[30] Methylotrophs provide plants with nutrients such as soluble phosphorus and fixed nitrogen and also play a role in the uptake of said nutrients.[29][30] Additionally, they can help plants respond to environmental stressors through the production of phytohormones.[29] Methylotrophic growth also inhibits the growth of harmful plant pathogens and induces systemic resistance.[30] Methylotrophic biofertilizers used either alone or together with chemical fertilizers have been shown to increase both crop yield and quality without loss of nutrients.[29]
References
- Anthony, C. "The Biochemistry of Methylotrophs". Academic press, 1982, p. 2-3
- Yurimoto, Hiroya; Kato, Nobuo; Sakai, Yasuyoshi (2005-01-01). "Assimilation, dissimilation, and detoxification of formaldehyde, a central metabolic intermediate of methylotrophic metabolism". The Chemical Record. 5 (6): 367–375. doi:10.1002/tcr.20056. ISSN 1528-0691. PMID 16278835.
- Nguyen, Ngoc-Loi; Yu, Woon-Jong; Yang, Hye-Young; Kim, Jong-Geol; Jung, Man-Young; Park, Soo-Je; Roh, Seong-Woon; Rhee, Sung-Keun (28 September 2017). "A novel methanotroph in the genus Methylomonas that contains a distinct clade of soluble methane monooxygenase". Journal of Microbiology. 55 (10): 775–782. doi:10.1007/s12275-017-7317-3. PMID 28956349.
- Ross, Matthew O.; Rosenzweig, Amy C. (2017-04-01). "A tale of two methane monooxygenases". JBIC Journal of Biological Inorganic Chemistry. 22 (2–3): 307–319. doi:10.1007/s00775-016-1419-y. ISSN 0949-8257. PMC 5352483. PMID 27878395.
- Hanson, R. S.; Hanson, T. E. (1996-06-01). "Methanotrophic bacteria". Microbiological Reviews. 60 (2): 439–471. ISSN 1092-2172. PMC 239451. PMID 8801441.
- Vorholt, Julia A. (2002-10-01). "Cofactor-dependent pathways of formaldehyde oxidation in methylotrophic bacteria". Archives of Microbiology. 178 (4): 239–249. doi:10.1007/s00203-002-0450-2. ISSN 0302-8933. PMID 12209256.
- J Colby; H Dalton; Whittenbury, and R. (1979). "Biological and Biochemical Aspects of Microbial Growth on C1 Compounds". Annual Review of Microbiology. 33 (1): 481–517. doi:10.1146/annurev.mi.33.100179.002405. PMID 386931.
- Oremland, Ronald S.; Kiene, Ronald P.; Mathrani, Indra; Whiticar, Michael J.; Boone, David R. (1989-04-01). "Description of an Estuarine Methylotrophic Methanogen Which Grows on Dimethyl Sulfide". Applied and Environmental Microbiology. 55 (4): 994–1002. ISSN 0099-2240. PMC 184236. PMID 16347900.
- Holmes, Andrew J.; Kelly, D. P.; Baker, Simon C.; Thompson, A. S.; Marco, Paolo De; Kenna, Elizabeth M.; Murrell, J. Colin (1997-01-01). "Methylosulfonomonas methylovora gen. nov., sp. nov., and Marinosulfonomonas methylotropha gen. nov., sp. nov.: novel methylotrophs able to grow on methanesulfonic acid". Archives of Microbiology. 167 (1): 46–53. doi:10.1007/s002030050415. ISSN 0302-8933. PMID 9000341.
- Kelly, Don P.; Baker, Simon C.; Trickett, Jim; Davey, Margaret; Murrell, J. Colin (1994). "Methanesulphonate utilization by a novel methylotrophic bacterium involves an unusual monooxygenase". Microbiology. 140 (6): 1419–1426. doi:10.1099/00221287-140-6-1419.
- Firsova, Julia; Doronina, Nina; Lang, Elke; Spröer, Cathrin; Vuilleumier, Stéphane; Trotsenko, Yuri (2009). "Ancylobacter dichloromethanicus sp. nov. – a new aerobic facultatively methylotrophic bacterium utilizing dichloromethane". Systematic and Applied Microbiology. 32 (4): 227–232. doi:10.1016/j.syapm.2009.02.002. PMID 19346095.
- Duine, J.a.; Frank, J.; Berkhout, M.p.j. (1984-03-26). "NAD-dependent, PQQ-containing methanol dehydrogenase: a bacterial dehydrogenase in a multienzyme complex". FEBS Letters. 168 (2): 217–221. doi:10.1016/0014-5793(84)80249-5. ISSN 1873-3468. PMID 6373362.
- Murray, William D.; Duff, Sheldon J. B.; Lanthier, Patricia H. (1989-11-01). "Induction and stability of alcohol oxidase in the methylotrophic yeast Pichia pastoris". Applied Microbiology and Biotechnology. 32 (1): 95–100. doi:10.1007/BF00164829. ISSN 0175-7598.
- Verseveld, H. W. Van; Stouthamer, A. H. (1978-07-01). "Electron-transport chain and coupled oxidative phosphorylation in methanol-grown Paracoccus denitrificans". Archives of Microbiology. 118 (1): 13–20. doi:10.1007/BF00406068. ISSN 0302-8933. PMID 29587.
- Chistoserdova, Ludmila; Crowther, Gregory J.; Vorholt, Julia A.; Skovran, Elizabeth; Portais, Jean-Charles; Lidstrom, Mary E. (2007-12-15). "Identification of a Fourth Formate Dehydrogenase in Methylobacterium extorquens AM1 and Confirmation of the Essential Role of Formate Oxidation in Methylotrophy". Journal of Bacteriology. 189 (24): 9076–9081. doi:10.1128/jb.01229-07. ISSN 0021-9193. PMC 2168636. PMID 17921299.
- Reed, William M.; Dugan, Patrick R. (1987). "Isolation and Characterization of the Facultative Methylotroph Mycobacterium ID-Y". Microbiology. 133 (5): 1389–1395. doi:10.1099/00221287-133-5-1389. PMID 3655742.
- van der Klei, Ida J.; Yurimoto, Hiroya; Sakai, Yasuyoshi; Veenhuis, Marten (2006). "The significance of peroxisomes in methanol metabolism in methylotrophic yeast". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1763 (12): 1453–1462. doi:10.1016/j.bbamcr.2006.07.016. PMID 17023065.
- TAYLOR, STEPHEN C.; DALTON, HOWARD; DOW, CRAWFORD S. (1981). "Ribulose-1,5-bisphosphate Carboxylase/Oxygenase and Carbon Assimilation in Methylococcus capsulatus (Bath)". Microbiology. 122 (1): 89–94. doi:10.1099/00221287-122-1-89.
- Raines, Christine A. (2003-01-01). "The Calvin cycle revisited". Photosynthesis Research. 75 (1): 1–10. doi:10.1023/A:1022421515027. ISSN 0166-8595. PMID 16245089.
- Champigny, Marie-Louise; Bismuth, Evelyne (1976-01-01). "Role of Photosynthetic Electron Transfer in Light Activation of Calvin Cycle Enzymes". Physiologia Plantarum. 36 (1): 95–100. doi:10.1111/j.1399-3054.1976.tb05034.x. ISSN 1399-3054.
- Taylor, Stephen; Dalton, Howard; Dow, Crawford (1980-07-01). "Purification and initial characterisation of ribulose 1,5-bisphosphate carboxylase from Methylococcus capsulatus (Bath)". FEMS Microbiology Letters. 8 (3): 157–160. doi:10.1016/0378-1097(80)90021-x.
- Quayle, J. R. (1980-02-01). "Microbial assimilation of C1 compounds". Biochemical Society Transactions. 8 (1): 1–10. doi:10.1042/bst0080001. ISSN 0300-5127. PMID 6768606.
- Chistoserdova, L. V.; Lidstrom, M. E. (1994-12-01). "Genetics of the serine cycle in Methylobacterium extorquens AM1: identification, sequence, and mutation of three new genes involved in C1 assimilation, orf4, mtkA, and mtkB". Journal of Bacteriology. 176 (23): 7398–7404. doi:10.1128/jb.176.23.7398-7404.1994. ISSN 0021-9193. PMC 197134. PMID 7961516.
- Yurimoto, Hiroya; Oku, Masahide; Sakai, Yasuyoshi (2011). "Yeast Methylotrophy: Metabolism, Gene Regulation and Peroxisome Homeostasis". International Journal of Microbiology. 2011: 101298. doi:10.1155/2011/101298. ISSN 1687-918X. PMC 3132611. PMID 21754936.
- Gonchar, Mykhailo; Maidan, Mykola; Korpan, Yaroslav; Sibirny, Volodymyr; Kotylak, Zbigniew; Sibirny, Andrei (2002-08-01). "Metabolically engineered methylotrophic yeast cells and enzymes as sensor biorecognition elements". FEMS Yeast Research. 2 (3): 307–314. doi:10.1016/s1567-1356(02)00124-1. ISSN 1567-1356. PMID 12702280.
- Gleeson, M. A.; Sudbery, P. E. (1988-03-01). "The methylotrophic yeasts". Yeast. 4 (1): 1–15. doi:10.1002/yea.320040102. ISSN 1097-0061.
- WAITES, M. J.; QUAYLE, J. R. (1981). "The Interrelation between Transketolase and Dihydroxyacetone Synthase Activities in the Methylotrophic Yeast Candida boidinii". Microbiology. 124 (2): 309–316. doi:10.1099/00221287-124-2-309. PMID 6276498.
- Guillaume Borrel, Didier Jézéquel, Corinne Biderre-Petit, Nicole Morel-Desrosiers, Jean-Pierre Morel, Pierre Peyret, Gérard Fonty, Anne-Catherine Lehours, Production and consumption of methane in freshwater lake ecosystems, In Research in Microbiology, Volume 162, Issue 9, 2011, Pages 832-847, ISSN 0923-2508, https://doi.org/10.1016/j.resmic.2011.06.004.
- Kumar, Manish; Singh Tomar, Rajesh; Lade, Harshad; Paul, Diby (2016). "Methylotrophic bacteria in sustainable agriculture". World Journal of Microbiology and Biotechnology. 32 (7): 120. doi:10.1007/s11274-016-2074-8. PMID 27263015.
- Iguchi, H.; Yurimoto, H.; Sakai, Y. (2015). "Interactions of methylotrophs with plants and other heterotrophic bacteria". Microorganisms. 3 (2): 137–151. doi:10.3390/microorganisms3020137. PMC 5023238. PMID 27682083.