Scaffold/matrix attachment region
The term S/MAR (scaffold/matrix attachment region), otherwise called SAR (scaffold-attachment region), or MAR (matrix-associated region), are sequences in the DNA of eukaryotic chromosomes where the nuclear matrix attaches. As architectural DNA components that organize the genome of eukaryotes into functional units within the cell nucleus, S/MARs mediate structural organization of the chromatin within the nucleus. These elements constitute anchor points of the DNA for the chromatin scaffold and serve to organize the chromatin into structural domains. Studies on individual genes led to the conclusion that the dynamic and complex organization of the chromatin mediated by S/MAR elements plays an important role in the regulation of gene expression.
Overview
It has been known for many years that a polymer meshwork, a so-called "nuclear matrix" or "nuclear-scaffold" is an essential component of eukaryotic nuclei. This nuclear skeleton acts as a dynamic support for many specialized events concerning the readout a spread of genetic information (see below).
S/MARs map to non-random locations in the genome. They occur at the flanks of transcribed regions, in 5´-introns, and also at gene breakpoint cluster regions (BCRs). Being association points for common nuclear structural proteins S/MARs are required for authentic and efficient chromosomal replication and transcription, for recombination and chromosome condensation. S/MARs do not have an obvious consensus sequence. Although prototype elements consist of AT-rich regions several hundred base pairs in length, the overall base composition is definitely not the primary determinant of their activity. Instead, their function requires a pattern of "AT-patches" that confer the propensity for local strand unpairing under torsional strain.
Bioinformatics approaches support the idea that, by these properties, S/MARs not only separate a given transcriptional unit (chromatin domain) from its neighbors, but also provide platforms for the assembly of factors enabling transcriptional events within a given domain. An increased propensity to separate the DNA strands (the so-called 'stress induced duplex destabilization' potential, SIDD) can serve the formation of secondary structures such as cruciforms or slippage structures, which are recognizable features for a number of enzymes (DNAses, topoisomerases, poly(ADP-ribosyl) polymerases and enzymes of the histone-acetylation and DNA-methylation apparatus). S/MARs have been classified as either being constitutive (acting as permanent domain boundaries in all cell types) or facultative (cell type- and activity-related) depending on their dynamic properties.
While the number of S/MARs in the human genome has been estimated to approach 64,000 (chromatin domains) plus an additional 10,000 (replication foci), in 2007 still only a minor fraction (559 for all eukaryotes) had met the standard criteria for an annotation in the S/MARt database SMARtDB.
Context-dependent properties
Current views of the nuclear matrix envision it as a dynamic entity, which changes its properties along the requirements of the cell nucleus—much the same as the cytoskeleton adapts its structure and function to external signals. In retrospect it is of note that the discovery of S/MARs has two major routes:
- the description of scaffold-attachment elements (SARs) by Laemmli and coworkers, which were thought to demarcate the borders of a given chromatin domain[1]
- the characterization of matrix-associated regions (MARs) the first examples of which supported the immunoglobulin kapp-chain enhancer according to its occupancy with transcription factors[2]
Subsequent work demonstrated both the constitutive (SAR-like) and the facultative (MAR-like) function of the elements depending on the context. Whereas constitutive S/MARs were found to be associated with a DNase I hypersensitive site in 'all' cell types (whether or not the enclosed domain was transcribed), DNAse I hypersensitivity of the facultative type depended on the transcriptional status.[3] The major difference between these two functional types of S/MARs is their size: the constitutive elements may extend over several kilobasepairs whereas facultative ones are at the lower size limit around 300 base pairs.
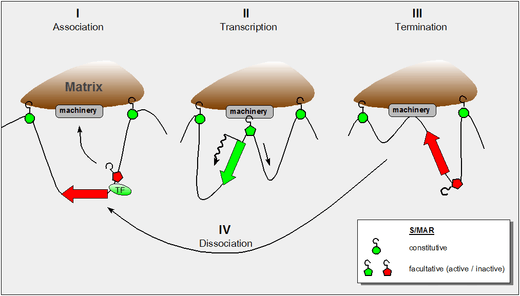
The figure shows our present understanding of these properties and it incorporates the following findings:
- the dynamic properties of S/MAR-scaffold contacts as derived by haloFISH investigations[4]
- the fact that during transcription DNA is reeled through RNA-polymerase which itself is a fixed component of the nuclear matrix[5]
- the fact that certain domain-intrinsic S/MARs require the support of an adjacent transcription factor to become active.[3]
Additional information
Recently, Tetko has found a strong correlation of intragenic S/MARs with spatiotemporal expression of genes in Arabidopsis thaliana.[6] On a genome scale, pronounced tissue- and organ-specific and developmental expression patterns of S/MAR-containing genes have been detected. Notably, transcription factor genes contain a significant higher portion of S/MARs. The pronounced difference in expression characteristics of S/MAR-containing genes emphasizes their functional importance and the importance of structural chromosomal characteristics for gene regulation in plants as well as within other eukaryotes.
References
- Mirkovitch J, Mirault ME, Laemmli UK (November 1984). "Organization of the higher-order chromatin loop: specific DNA attachment sites on nuclear scaffold". Cell. 39 (1): 223–32. doi:10.1016/0092-8674(84)90208-3. PMID 6091913.
- Cockerill PN, Garrard WT (January 1986). "Chromosomal loop anchorage of the kappa immunoglobulin gene occurs next to the enhancer in a region containing topoisomerase II sites". Cell. 44 (2): 273–82. doi:10.1016/0092-8674(86)90761-0. PMID 3002631.
- Klar M, Stellamanns E, Ak P, Gluch A, Bode J (December 2005). "Dominant genomic structures: detection and potential signal functions in the interferon-beta domain". Gene. 364: 79–89. doi:10.1016/j.gene.2005.07.023. PMID 16185826.
- Heng HH, Goetze S, Ye CJ, et al. (March 2004). "Chromatin loops are selectively anchored using scaffold/matrix-attachment regions". J. Cell Sci. 117 (Pt 7): 999–1008. doi:10.1242/jcs.00976. PMID 14996931.
- Jackson DA, Dolle A, Robertson G, Cook PR (August 1992). "The attachments of chromatin loops to the nucleoskeleton". Cell Biol. Int. Rep. 16 (8): 687–96. doi:10.1016/s0309-1651(05)80013-x. PMID 1446346.
- See Tetko, Igor V., Georg Haberer, Stephen Rudd, Blake Meyers, Hans-Werner Mewes, and Klaus F. X. Mayer (2006) Spatiotemporal Expression Control Correlates with Intragenic Scaffold Matrix Attachment Regions (S/MARs) in Arabidopsis thaliana. PLoS Computational Biology 2:136-145. Online copy.