Clostridioides difficile (bacteria)
Clostridioides difficile (syn. Clostridium difficile), also known as Peptoclostridium difficile, C. difficile, or C. diff (/siː dɪf/), is Gram-positive species of spore-forming bacteria.[3]
| Clostridioides difficile | |
|---|---|
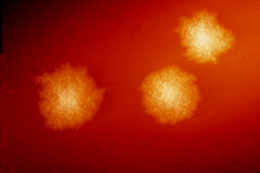 | |
| C. difficile colonies on a blood agar plate | |
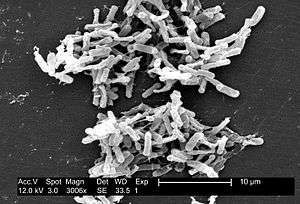 | |
| Micrograph of Clostridioides difficile | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Firmicutes |
| Class: | Clostridia |
| Order: | Clostridiales |
| Family: | Peptostreptococcaceae |
| Genus: | Clostridioides |
| Species: | C. difficile |
| Binomial name | |
| Clostridioides difficile (Hall & O'Toole, 1935) Lawson & Rainey, 2016 | |
| Synonyms | |
Clostridioides spp. are anaerobic, motile bacteria, ubiquitous in nature, and especially prevalent in soil. Its vegetative cells are rod-shaped, pleomorphic, and occur in pairs or short chains. Under the microscope, they appear as long, irregular (often drumstick- or spindle-shaped) cells with a bulge at their terminal ends (forms subterminal spores). Under Gram staining, C. difficile cells are Gram-positive and show optimum growth on blood agar at human body temperatures in the absence of oxygen. C. difficile is catalase- and superoxide dismutase-negative, and produces two types of toxins: enterotoxin A and cytotoxin B, which disrupts cytoskeleton signal transductions in the host.[4] Under stress conditions, the bacteria produce spores that are able to tolerate extreme conditions that the active bacteria cannot tolerate.[5]
C. difficile may become established in the human colon; it is present in 2–5% of the adult population.[5] Sometimes antibiotic therapy for various infections has the adverse effect of disrupting the normal balance of the gut microbiota, in which case C. difficile may opportunistically dominate, causing C. difficile infection (CDI).
Taxonomy
The species was transferred from the genus Clostridium to Clostridioides in 2016, thus giving it the binomial Clostridioides difficile.[6][7][8] This new name reflects the taxonomic differences between this species and other members of the genus Clostridium, while maintaining the common name as C. diff.[9] As of 2018, the only other species in this new genus Clostridioides mangenotii (formerly known as Clostridium mangenotii).[10] A July 2013 paper from Environmental Microbiology proposed to rename the species Peptoclostridium difficile.[11][12]
Human pathogen
Pathogenic C. difficile strains produce multiple toxins.[13] The best-characterized are enterotoxin (C. difficile toxin A) and cytotoxin (C. difficile toxin B), both of which may produce diarrhea and inflammation in infected patients (C. difficile colitis), although their relative contributions have been debated. The diarrhea may range from a few days of intestinal fluid loss to life-threatening pseudomembranous colitis, which is associated with intense inflammation of the colon and formation of pseudomembranes on the intestinal mucosal surface.[5] Toxins A and B are glucosyltransferases that target and inactivate the Rho family of GTPases. Toxin B (cytotoxin) induces actin depolymerization by a mechanism correlated with a decrease in the ADP-ribosylation of the low molecular mass GTP-binding Rho proteins.[14] There is also a binary toxin (AB toxin), but its role in disease is not fully understood.[15]
Additional virulence factors include an adhesin factor that mediates the binding to human colonic cells and a hyaluronidase.[16] The bacterium also produces the chemical para-cresol, which inhibits the growth of other microbes in its vicinity and allows it to outcompete normal human gut flora.[17]
Antibiotic treatment of C. diff infections may be difficult, due both to antibiotic resistance and physiological factors of the bacterium (spore formation, protective effects of the pseudomembrane).[5] The emergence of a new, highly toxic strain of C. difficile, resistant to fluoroquinolone antibiotics, such as ciprofloxacin and levofloxacin, said to be causing geographically dispersed outbreaks in North America, was reported in 2005.[18] The U.S. Centers for Disease Control in Atlanta warned of the emergence of an epidemic strain with increased virulence, antibiotic resistance, or both.[19] Resistance to other antibiotics such as metronidazole, the first choice of antimicrobial drug when treating CDI, has been observed in up to 12% of clinical isolates, so as treatment with various antibiotics continues, more diverse and stronger resistances will continue to evolve in C. difficile populations, further complicating attempts at effective treatment.[20]
Transmission
C. difficile is transmitted from person to person by the fecal-oral route, shed in faeces. Any surface, device, or material (e.g., toilets, bathing tubs, and electronic rectal thermometers) that becomes contaminated with faeces may serve as a reservoir for the C. difficile spores. C. difficile spores are transferred to patients mainly by the hands of healthcare personnel who have touched a contaminated surface or item. C. difficile can live for long periods of time on surfaces.[21] The organism forms heat-resistant spores that are not killed by alcohol-based hand cleansers or routine surface cleaning, thus, these spores survive in clinical environments for long periods. Because of this, the bacterium may be cultured from almost any surface. Once spores are ingested, their acid resistance allows them to pass through the stomach unscathed. They germinate and multiply into vegetative cells in the colon upon exposure to bile acids. Consequently, the World Health Organization advocates the use of soap in addition to alcohol solutions to limit the spread of the spores.[22] Sporulation was shown to be significantly reduced after inactivation of C. diffiicile's DNA methyltransferase CamA[23], raising the prospect of developing a drug that may inhibit this bacterium in a specific manner.
Host range
C. difficile infects pigs, calves, and humans, and inhabits a natural reservoir of soil, faeces of domestic animals and humans, sewage, the human intestinal tract, and retail meat.[24]
A 2015 CDC study estimated that C. diff afflicted almost half a million Americans and caused 29,000 deaths in 2011. The study estimated that 40% of cases began in nursing homes or community health-care settings, while 24% occurred in hospitals.[25]
C. difficile is common in the human digestive system. However, it is a poor competitor, and is often outcompeted for nutrients by other bacteria in the digestive system. As a result, C. difficile is kept to a manageable number. If the sudden introduction of antibiotic disrupts the microbiome, C. difficile may be able to grow as a result of many of its competitors being killed off. The incubation period is 5–10 days, with a range of 1 day to weeks following antibiotic treatment for antibiotic associated diarrhea. Additionally, carriage of C. difficile with high levels of toxins is common in young children, while disease is rare. The production of one or even both toxins is not always sufficient for producing symptoms.[26]
Signs and symptoms
Symptoms of C. difficile infection include: diarrhea (at least three loose bowel movements a day), dehydration, abdominal pain that can be severe, loss of appetite, and nausea.[27]
Treatment
Patients being treated with antibiotics when symptoms begin should stop taking them, if possible. This break in antibiotic therapy can sometimes lead to spontaneous resolution of symptoms. Patients who do not respond to the cessation of broad-spectrum antibiotics will need to be treated with antibiotics capable of killing C. difficile spores. Primary infections are typically treated with vancomycin, with a usual dosage of 125 mg every 6 hours.[28] The vancomycin regimen has replaced the traditional use of metronidazole due to its greater efficacy, safety profile, and lower recurrence rates. In patients who cannot tolerate vancomycin, fidaxomicin is an acceptable option with similar efficacy and even lower recurrence rates than vancomycin.[29] In cases of fulminant CDI, adjuvant therapy with parenteral metronizadole plus oral vancomycin or fidaxomicin is suggested.[30]
About 20% of patients who successfully complete therapy of primary infection with metronidazole or vancomycin will experience a relapse. A fraction of those patients will experience continuous reoccurrences of the infection. The first relapse of C. difficile is usually treated with the same antibiotic used to treat the primary infection. Any subsequent infections should not be treated with metronidazole. Occasionally, a standard 10-day course of oral vancomycin will not work. In these cases, a vancomycin taper is the preferred treatment. Patients take decreasing doses of vancomycin over a period of up to 3 months, depending on the severity of the infection.[27]
Each subsequent relapse of C. difficile tends to be more severe than previous infections. Long-term treatment with a vancomycin taper supplemented with probiotics, especially Saccharomyces boulardii, is associated with a higher rate of success.[31]
After three relapses, patients may be treated with oral fidaxomicin, a narrow-spectrum antibiotic. The usual dosage is 200 mg twice a day orally for 10 days. Fidaxomicin is considered to be superior to vancomycin for severe CDI.[32] The major downside of treatment with fidaxomicin is the cost of medication. A 10-day course may cost up to US$3500.
Patients who do not respond to traditional antibiotic therapy may be eligible for a Fecal microbiota transplant (FMT). Healthcare providers can transfer stool from a healthy person to the colon of a patient with repeated CDI. This process is the most successful treatment for severe CDI with a cure rate around 93%. Recurrence rates of CDI in patients treated with a FMT are generally low, around 19%, which makes it very effective at treating chronic CDI cases. However, in some cases, flares of inflammatory bowel disease are a possible side effect of the treatment.[33] Long-term effects of FMT are unknown, as the procedure has only been FDA approved since 2011 and relatively few procedures have been performed. If transplantation is not an option, removal of the infected part of the colon can cure CDI.[32][27]
Strains
In 2005, molecular analysis led to the identification of the C. difficile strain type characterized as group BI by restriction endonuclease analysis, as North American pulse-field-type NAP1 by pulsed-field gel electrophoresis and as ribotype 027; the differing terminology reflects the predominant techniques used for epidemiological typing. This strain is referred to as C. difficile BI/NAP1/027.[34]
As of 2016, the NAP1 strain has been replaced by novel strains in some areas of British Columbia. These novel strains include NAP2 and NAP4, and some strains that do not have a NAP designation. The frequency of these novel strains increased from 2008 to 2013 in one studied region, displacing the originally more common and recognizable NAP1 bacteria.[35]
Two strains, ribotypes RT078 and RT027, can live on low concentrations of the sugar trehalose; both strains became more common after trehalose was introduced as a food additive in the early 2000s, thus increasing dietary trehalose intake.[36]
Genome
| NCBI genome ID | 535 |
|---|---|
| Ploidy | haploid |
| Genome size | 4.3 Mb |
| Number of chromosomes | 1 |
| Year of completion | 2005 |
The first complete genome sequence of a C. difficile strain was first published in 2005 by Sanger Institute in the UK. This was of the strain 630, a virulent and multiple drug-resistant strain isolated in Switzerland in 1982. Scientists at Sanger Institute have sequenced genomes of about 30 C. difficile isolates using next-generation sequencing technologies from 454 Life Sciences and Illumina.[37]
Researchers at McGill University in Montreal sequenced the genome of the highly virulent Quebec strain of C. difficile in 2005 using ultra-high throughput sequencing technology. The tests involved doing 400,000 DNA parallel-sequencing reactions of the bacterium's genome, which had been fragmented for sequencing. These sequences were assembled computationally to form a complete genome sequence.[18][38]
In 2012, scientists at University of Oxford sequenced C. difficile genomes from 486 cases arising over four years in Oxfordshire using next-generation sequencing technologies from Illumina.[39]
Epigenome
C. difficile has a highly diverse epigenome, with 17 high-quality methylation motifs reported so far, the majority pertaining to the 6mA type. Methylation at one of these motifs - CAAAAA, was shown to impact sporulation, a key step in C. difficile disease transmission, as well as cell length, biofilm formation, and host colonization[23].
Bacteriophage
At least eight mainly temperate bacteriophages have been isolated from C. difficile, ranging in genome size from about 30 to about 60 kb.[40] Both environmentally and clinically derived C. difficile strains carry a diverse and prevalent set of prophages.[40]
Etymology and pronunciation
References
- Hall, Ivan C.; O'Toole, Elizabeth (1935). "Intestinal flora in new-born infants: with a description of a new pathogenic anaerobe, Bacillus difficilis". American Journal of Diseases of Children. 49 (2): 390–402. doi:10.1001/archpedi.1935.01970020105010.
- Prévot, A.-R. (1938). "Études de systématique bactérienne. IV. Critique de la conception actuelle du genre Clostridium". Annales de l'Institut Pasteur. 61 (1): 84.
- Moreno MA, Furtner F, Rivara FP (June 2013). "Clostridium difficile: A Cause of Diarrhea in Children". JAMA Pediatrics. 167 (6): 592. doi:10.1001/jamapediatrics.2013.2551. PMID 23733223.
- di Masi, Alessandra; Ascenzi, Paolo; Siarakas, Steven; Petrosillo, Nicola; Di Bella, Stefano (2016). "Clostridium difficile Toxins A and B: Insights into Pathogenic Properties and Extraintestinal Effects". Toxins (Basel). 8 (5): 134. doi:10.3390/toxins8050134. PMC 4885049. PMID 27153087.
- Ryan KJ, Ray CG, eds. (2004). Sherris Medical Microbiology (4th ed.). McGraw Hill. pp. 322–4. ISBN 978-0-8385-8529-0.
- Oren, Aharon; Garrity, George M. (2017). "List of new names and new combinations previously effectively, but not validly, published". International Journal of Systematic and Evolutionary Microbiology. 67 (9): 3140–3143. doi:10.1099/ijsem.0.002278. PMC 5817221. PMID 28891789.
- Lawson, Paul A.; Citron, Diane M.; Tyrrell, Kerin L.; Finegold, Sydney M. (August 2016). "Reclassification of Clostridium difficile as Clostridioides difficile (Hall and O'Toole 1935) Prévot 1938". Anaerobe. 40: 95–99. doi:10.1016/j.anaerobe.2016.06.008. ISSN 1095-8274. PMID 27370902.
- Zhu, Duolong; Sorg, Joseph A.; Sun, Xingmin (2018). "Clostridioides difficile Biology: Sporulation, Germination, and Corresponding Therapies for C. difficile Infection". Frontiers in Cellular and Infection Microbiology. 8: 29. doi:10.3389/fcimb.2018.00029. ISSN 2235-2988. PMC 5809512. PMID 29473021.
- Lawson, Paul A.; Citron, Diane M.; Tyrrell, Kerin L.; Finegold, Sydney M. (August 2016). "Reclassification of Clostridium difficile as Clostridioides difficile (Hall and O'Toole 1935) Prévot 1938". Anaerobe. 40: 95–99. doi:10.1016/j.anaerobe.2016.06.008. ISSN 1075-9964. PMID 27370902.
- Galperin, Michael Y.; Brover, Vyacheslav; Tolstoy, Igor; Yutin, Natalya (2016). "Phylogenomic analysis of the family Peptostreptococcaceae (Clostridium cluster XI) and proposal for reclassification of Clostridium litorale (Fendrich et al. 1991) and Eubacterium acidaminophilum (Zindel et al. 1989) as Peptoclostridium litorale gen. nov. comb. nov. and Peptoclostridium acidaminophilum comb. nov". International Journal of Systematic and Evolutionary Microbiology. 66 (12): 5506–5513. doi:10.1099/ijsem.0.001548. PMC 5244501. PMID 27902180.
- Yutin N, Galperin MY (2013). "A genomic update on clostridial phylogeny: Gram-negative spore formers and other misplaced clostridia". Environ. Microbiol. 15 (10): 2631–41. doi:10.1111/1462-2920.12173. PMC 4056668. PMID 23834245.
- Gerard J. Tortora; Berdell R. Funke; Christine L. Case (2015-01-13). Microbiology: An Introduction. Pearson Education. ISBN 978-0-13-392339-1.
- Di Bella, Stefano; Ascenzi, Paolo; Siarakas, Steven; Petrosillo, Nicola; di Masi, Alessandra (2016-01-01). "Clostridium difficile Toxins A and B: Insights into Pathogenic Properties and Extraintestinal Effects". Toxins. 8 (5): 134. doi:10.3390/toxins8050134. ISSN 2072-6651. PMC 4885049. PMID 27153087.
- Just I, Selzer J, von Eichel-Streiber C, Aktories K (1995). "The low molecular mass GTP-binding protein Rh is affected by toxin a from Clostridium difficile". The Journal of Clinical Investigation. 95 (3): 1026–31. doi:10.1172/JCI117747. PMC 441436. PMID 7883950.
- Barth H, Aktories K, Popoff MR, Stiles BG (2004). "Binary Bacterial Toxins: Biochemistry, Biology, and Applications of Common Clostridium and Bacillus Proteins". Microbiology and Molecular Biology Reviews. 68 (3): 373–402, table of contents. doi:10.1128/MMBR.68.3.373-402.2004. PMC 515256. PMID 15353562.
- [Medical Micriobiology, Fifth Edition, Patrick Murray, Elsevier Mosby, 2005, page 412]
- "The chemical weapon that helps bacterium wreak havoc in the gut". Nature: International Journal of Science. 14 September 2018. Retrieved 8 October 2018.
- Loo VG, Poirier L, Miller MA, Oughton M, Libman MD, Michaud S, Bourgault AM, Nguyen T, Frenette C, Kelly M, Vibien A, Brassard P, Fenn S, Dewar K, Hudson TJ, Horn R, René P, Monczak Y, Dascal A (December 2005). "A predominantly clonal multi-institutional outbreak of Clostridium difficile-associated diarrhea with high morbidity and mortality". The New England Journal of Medicine. 353 (23): 2442–9. doi:10.1056/NEJMoa051639. PMID 16322602.
- McDonald LC (August 2005). "Clostridium difficile: responding to a new threat from an old enemy" (PDF). Infection Control and Hospital Epidemiology. 26 (8): 672–5. doi:10.1086/502600. PMID 16156321.
- Saeed S. Banawas (21 February 2018). "Clostridium difficile Infections: A Global Overview of Drug Sensitivity and Resistance Mechanisms". Hindawi. BioMed Research International. Retrieved 4 November 2018.
- "Clostridium difficile Infection Information for Patients | HAI | CDC". www.cdc.gov. Retrieved 22 May 2017.
- "WHO Guidelines on Hand Hygiene in Health Care: a Summary" (PDF). World Health Organization. 2009. p. 31. Retrieved June 18, 2018.
- Oliveira, Pedro H.; Ribis, John W.; Garrett, Elizabeth M.; Trzilova, Dominika; Kim, Alex; Sekulovic, Ognjen; Mead, Edward A.; Pak, Theodore; Zhu, Shijia; Deikus, Gintaras; Touchon, Marie (2019-11-25). "Epigenomic characterization of Clostridioides difficile finds a conserved DNA methyltransferase that mediates sporulation and pathogenesis". Nature Microbiology: 1–15. doi:10.1038/s41564-019-0613-4. ISSN 2058-5276. PMID 31768029.
- Gould, L. Hannah; Limbago, Brandi (2010). "Clostridium difficile in Food and Domestic Animals: A New Foodborne Pathogen?". Clinical Infectious Diseases. 51 (5): 577–82. doi:10.1086/655692. PMID 20642351.
- Belluck, Pam (February 25, 2015). "Death Toll From C. Difficile Is Raised". The New York Times. Retrieved 25 February 2015.
- [Medical Microbiology, Fifth Edition, Patrick Murray, Elsevier Mosby, 2005, page 412]
- https://www.cdc.gov/hai/organisms/cdiff/cdiff-patient.html
- McDonald, L. Clifford; Gerding, Dale N.; Johnson, Stuart; Bakken, Johan S.; Carroll, Karen C.; Coffin, Susan E.; Dubberke, Erik R.; Garey, Kevin W.; Gould, Carolyn V. (2018-03-19). "Clinical Practice Guidelines for Clostridium difficile Infection in Adults and Children: 2017 Update by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA)". Clinical Infectious Diseases. 66 (7): e1–e48. doi:10.1093/cid/cix1085. ISSN 1537-6591. PMC 6018983. PMID 29462280.
- Mullane, Kathleen M.; Miller, Mark A.; Weiss, Karl; Lentnek, Arnold; Golan, Yoav; Sears, Pamela S.; Shue, Youe-Kong; Louie, Thomas J.; Gorbach, Sherwood L. (September 2011). "Efficacy of fidaxomicin versus vancomycin as therapy for Clostridium difficile infection in individuals taking concomitant antibiotics for other concurrent infections". Clinical Infectious Diseases. 53 (5): 440–447. doi:10.1093/cid/cir404. ISSN 1537-6591. PMC 3156139. PMID 21844027.
- Shane, Andi L.; Mody, Rajal K.; Crump, John A.; Tarr, Phillip I.; Steiner, Theodore S.; Kotloff, Karen; Langley, Joanne M.; Wanke, Christine; Warren, Cirle Alcantara (2017-11-29). "2017 Infectious Diseases Society of America Clinical Practice Guidelines for the Diagnosis and Management of Infectious Diarrhea". Clinical Infectious Diseases. 65 (12): e45–e80. doi:10.1093/cid/cix669. ISSN 1537-6591. PMC 5850553. PMID 29053792.
- "Treatment of Recurrent Clostridium difficile Colitis with Vancomycin and Saccharomyces boulardii" (PDF). The American Journal of Gastroenterology.
- Surawicz, Christina M; Brandt, Lawrence J; Binion, David G; Ananthakrishnan, Ashwin N; Curry, Scott R; Gilligan, Peter H; McFarland, Lynne V; Mellow, Mark; Zuckerbraun, Brian S (2013-02-26). "Guidelines for Diagnosis, Treatment and Prevention of Clostridium difficile Infections". The American Journal of Gastroenterology. 108 (4): 478–498. doi:10.1038/ajg.2013.4. ISSN 0002-9270. PMID 23439232.
- Chen, Yu-Gen; Zheng, Xiao; Zhou, Jin-Yong; Wu, Jing; Jiang, Feng; Zhang, Dan; Zhou, Qun; Chen, Tuo (2018-05-25). "Effect of Faecal Microbiota Transplantation for Treatment of Clostridium difficile Infection in Patients With Inflammatory Bowel Disease: A Systematic Review and Meta-Analysis of Cohort Studies". Journal of Crohn's and Colitis. 12 (6): 710–717. doi:10.1093/ecco-jcc/jjy031. ISSN 1873-9946. PMID 29528385.
- Rupnik M, Wilcox MH, Gerding DN (July 2009). "Clostridium difficile infection: New developments in epidemiology and pathogenesis". Nature Reviews. Microbiology. 7 (7): 526–36. doi:10.1038/nrmicro2164. PMID 19528959.
- Jassem, Agatha; Prystajecky, Natalie; Marra, Fawziah; Kibsey, Pamela; Tan, Kennard; Umlandt, Patricia; Janz, Loretta; Champagne, Sylvie; Gamage, Bruce; Golding, George; Mulvey, Michael; Henry, Bonnie; Hoang, Linda (29 Mar 2016). "Characterization of Clostridium difficile Strains in British Columbia, Canada: A Shift from NAP1 Majority (2008) to Novel Strain Types (2013) in One Region". Canadian Journal of Infectious Diseases and Medical Microbiology. 2016: 8207418. doi:10.1155/2016/8207418. PMC 4904575. PMID 27366181.
- Collins, J.; Robinson, C.; Danhof, H.; Knetsch, C. W.; van Leeuwen, H. C.; Lawley, T. D.; Auchtung, J. M.; Britton, R. A. (2018). "Dietary trehalose enhances virulence of epidemic Clostridium difficile". Nature. 553 (7688): 291–294. doi:10.1038/nature25178. ISSN 0028-0836. PMC 5984069. PMID 29310122.
- He M, Sebaihia M, Lawley TD, Stabler RA, Dawson LF, Martin MJ, Holt KE, Seth-Smith HM, Quail MA, Rance R, Brooks K, Churcher C, Harris D, Bentley SD, Burrows C, Clark L, Corton C, Murray V, Rose G, Thurston S, van Tonder A, Walker D, Wren BW, Dougan G, Parkhill J (April 2010). "Evolutionary dynamics of Clostridium difficile over short and long time scales". Proceedings of the National Academy of Sciences of the United States of America. 107 (16): 7527–32. Bibcode:2010PNAS..107.7527H. doi:10.1073/pnas.0914322107. PMC 2867753. PMID 20368420.
- Scientists map C. difficile strain – Institute of Public Affairs, Montreal
- Didelot X, Eyre DW, Cule M, Ip CL, Ansari MA, Griffiths D, Vaughan A, O'Connor L, Golubchik T, Batty EM, Piazza P, Wilson DJ, Bowden R, Donnelly PJ, Dingle KE, Wilcox M, Walker AS, Crook DW, A Peto TE, Harding RM (December 2012). "Microevolutionary analysis of Clostridium difficile genomes to investigate transmission" (PDF). Genome Biology. 13 (12): R118. doi:10.1186/gb-2012-13-12-r118. PMC 4056369. PMID 23259504.
- Hargreaves KR, Clokie MR (2014). "Clostridium difficile phages: Still difficult?". Frontiers in Microbiology. 5: 184. doi:10.3389/fmicb.2014.00184. PMC 4009436. PMID 24808893.
External links
- Pathogen Safety Data Sheets: Infectious Substances – Clostridium Difficile, Public Health Agency, Canada, 10 September 2014.
- Type strain of Clostridium difficile, BacDive - the Bacterial Diversity Metadatabase.